CMAdb
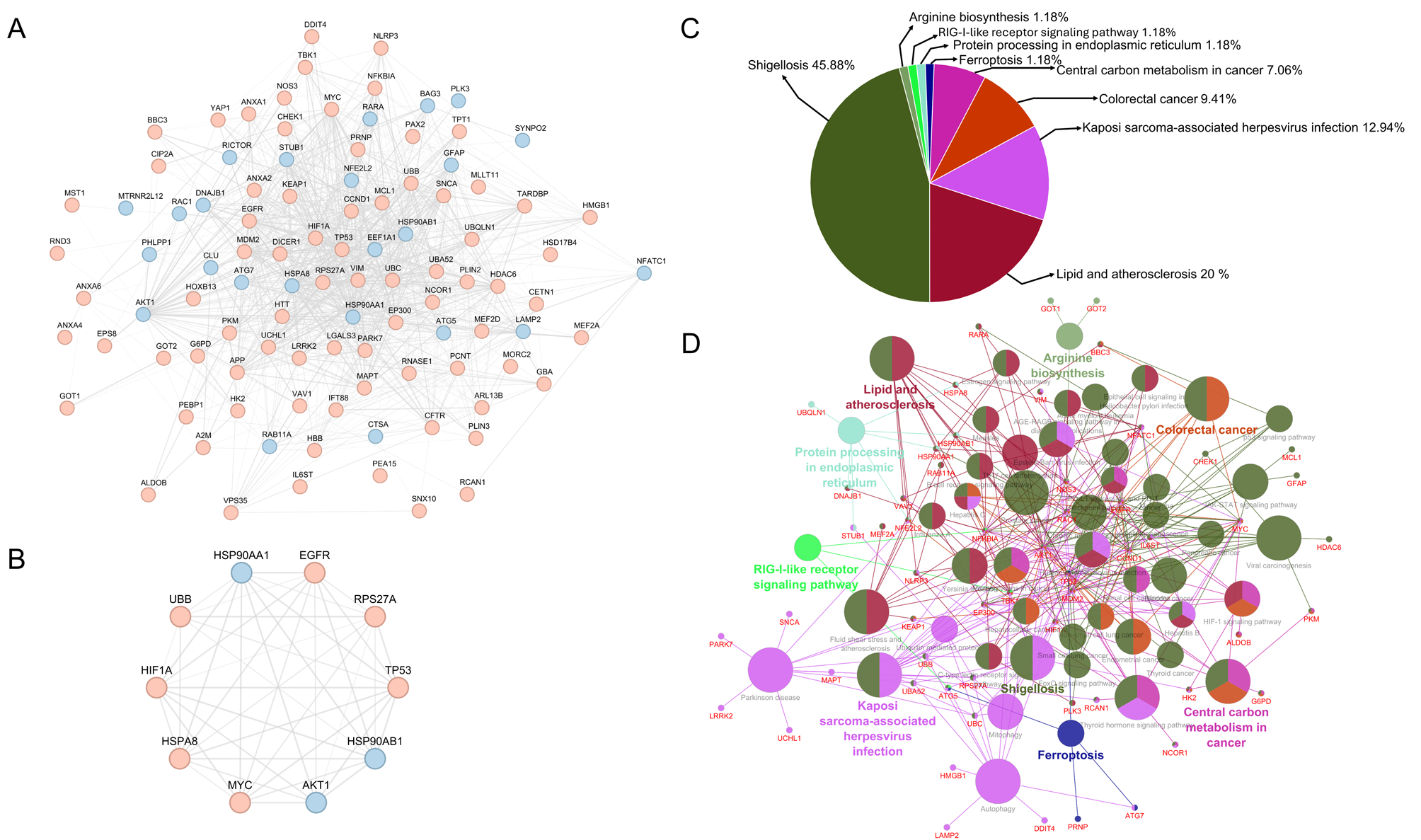
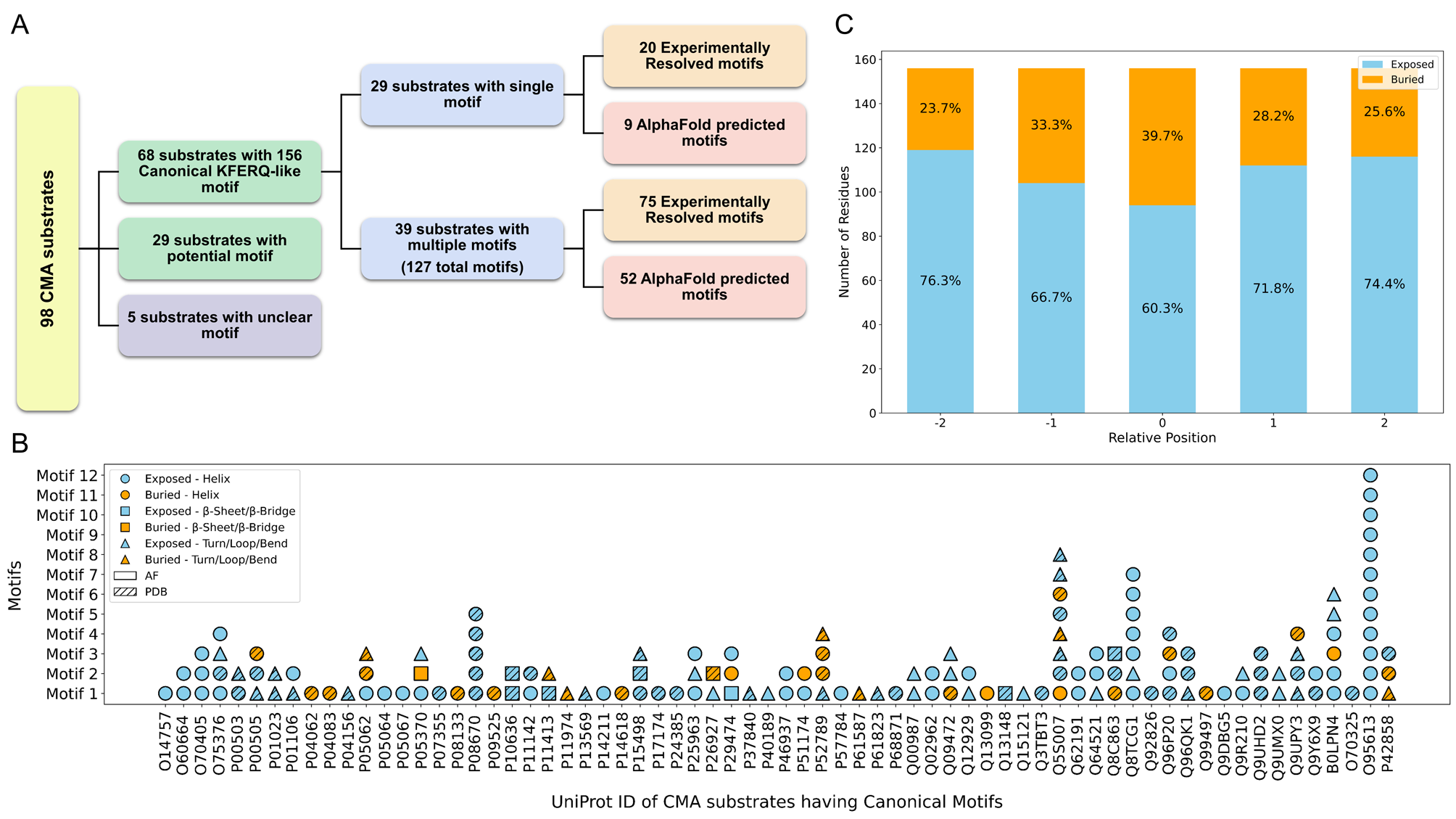
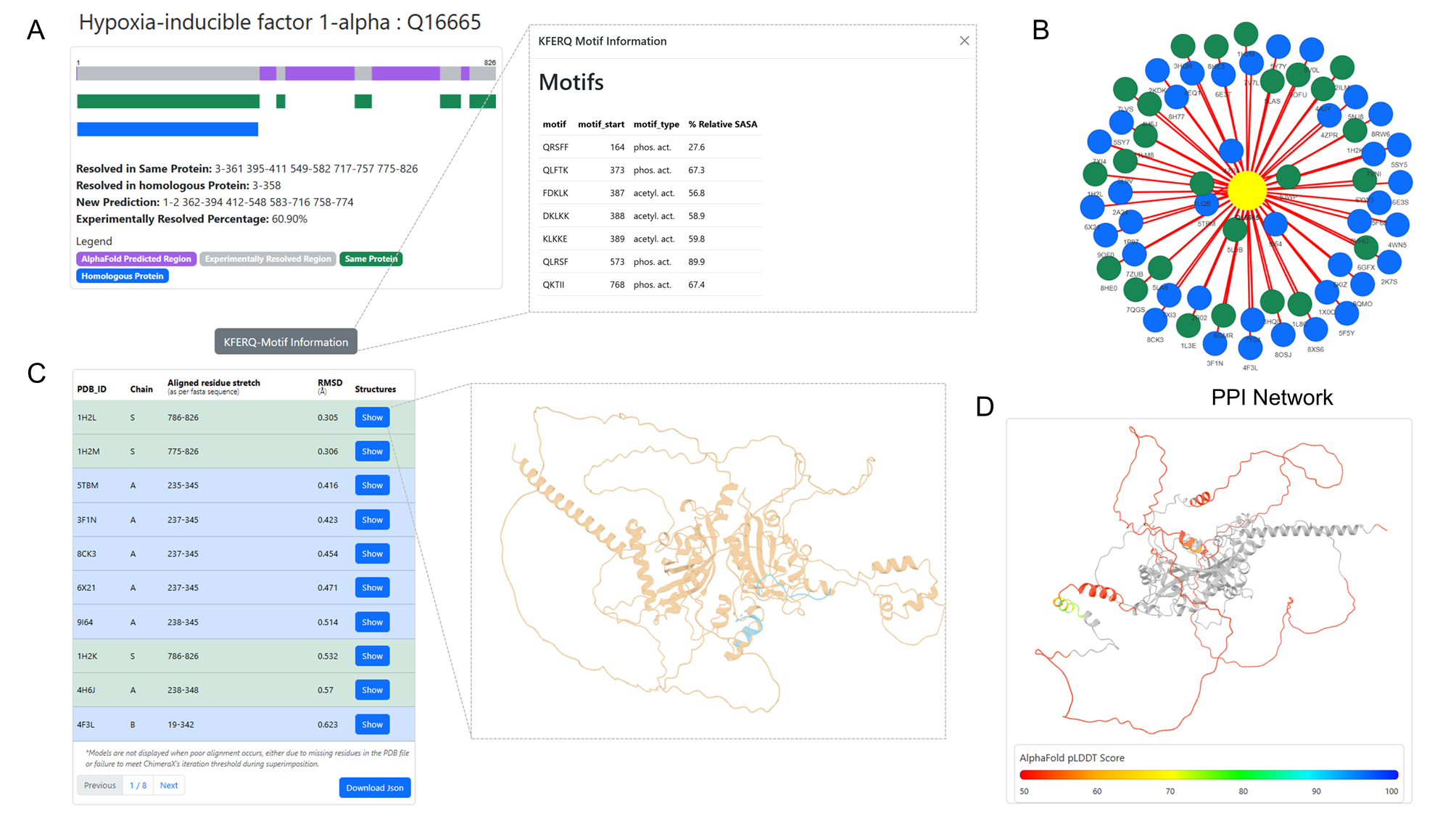
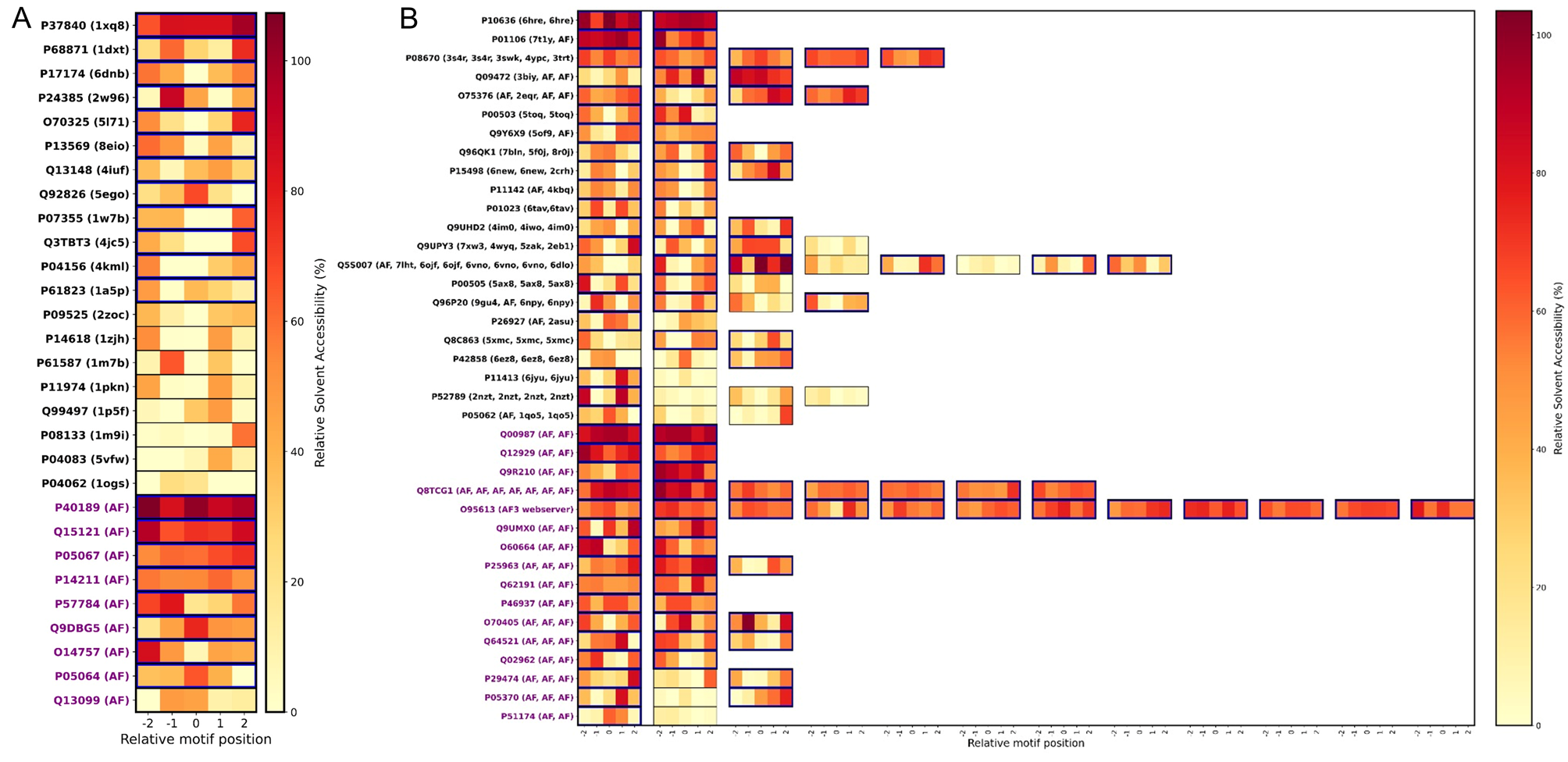
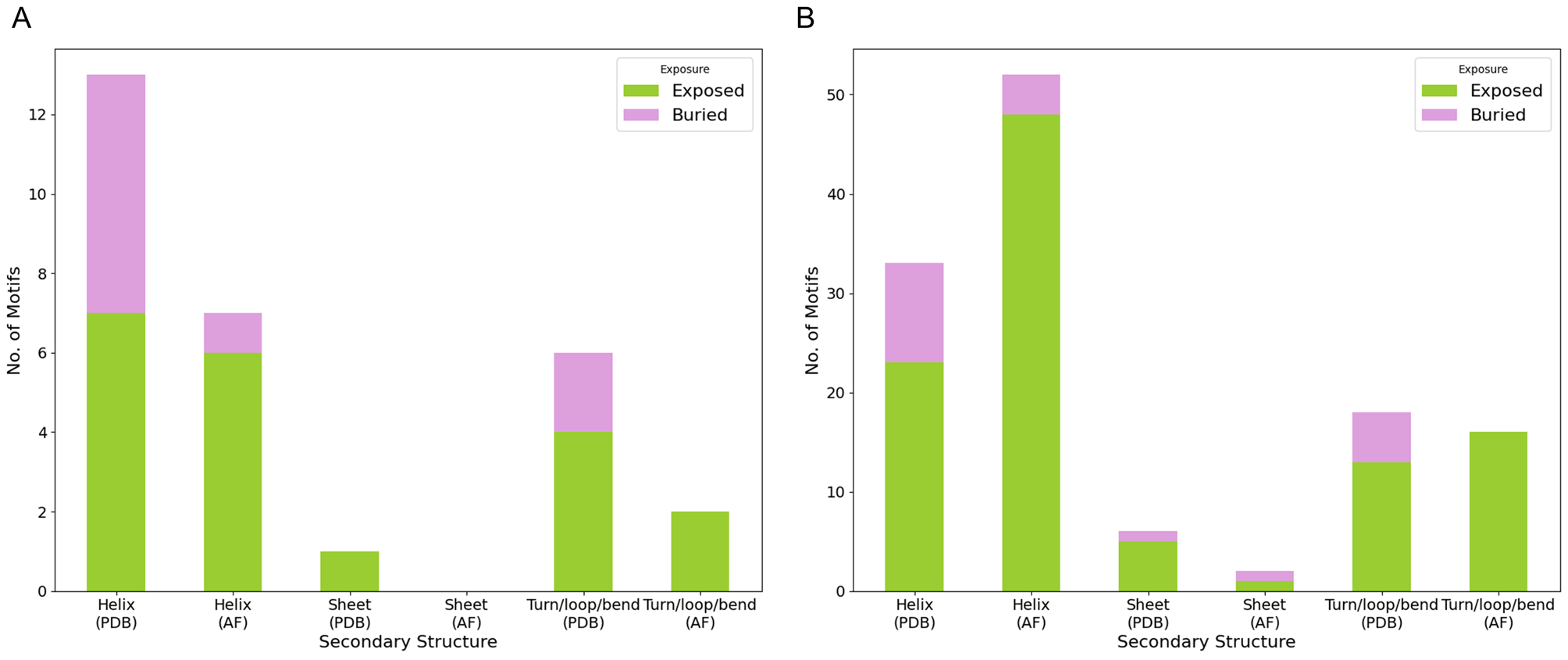
CMAdb is the first dedicated database that systematically catalogs experimentally-validated CMA substrates, their predicted domain information, mapping their ‘CMA-targeting’ motifs and calculating their solvent accessibility, structural alignments, and reliability scores. Our database serves as a foundational resource for researchers investigating the mechanistic underpinnings of CMA and its implications in health and disease.
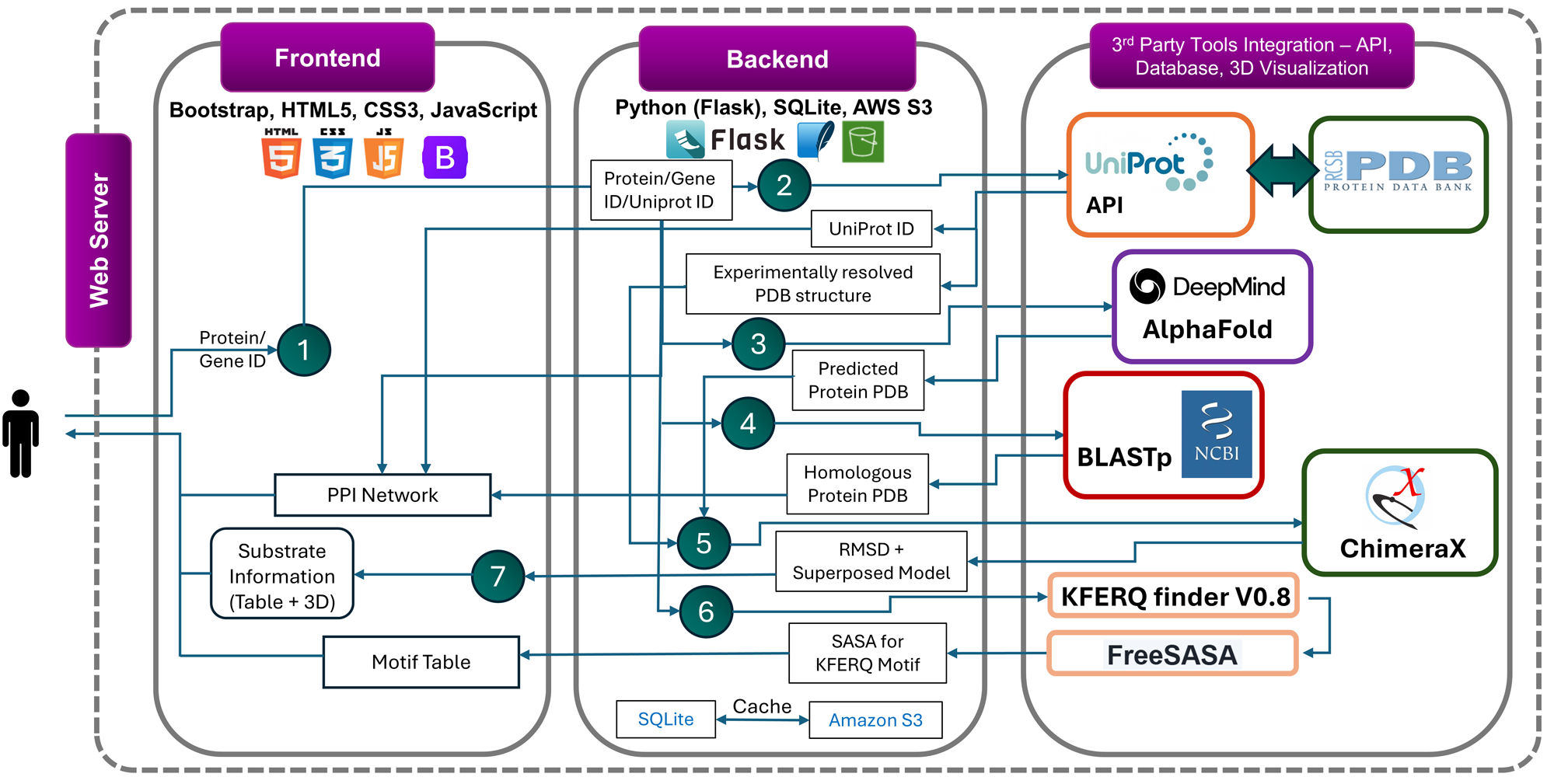
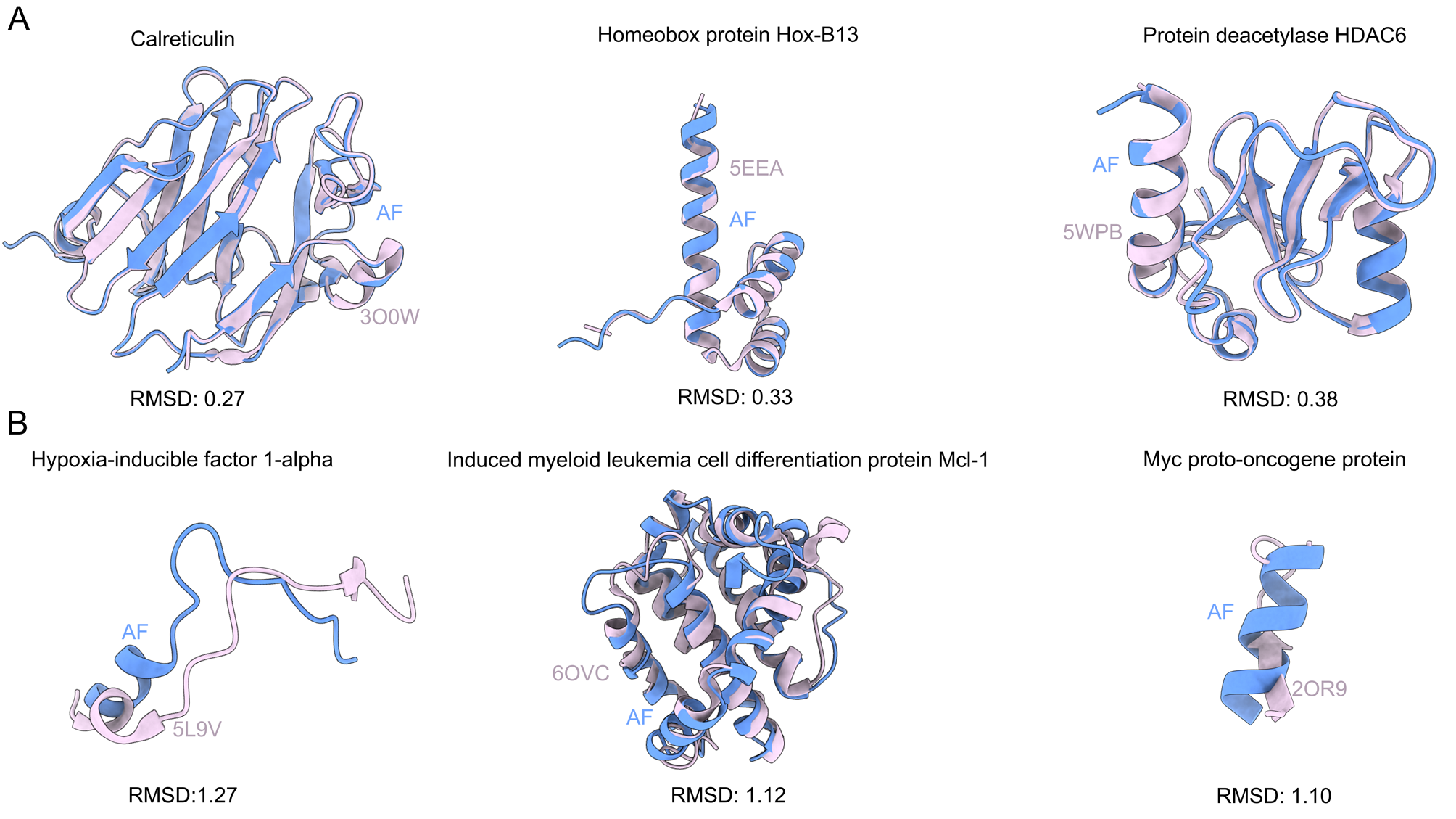
To support broader exploration, we developed an interactive web server that enables users to query proteins, retrieve all available PDB entries, identify homologous structures, perform structural superimposition with RMSD calculation, and highlight unresolved regions. For CMA substrates specifically, the server displays canonical and potential motif positions, and their solvent accessibility—providing a valuable platform for motif prediction and insights into CMA-mediated degradation.